Efficient, few-shot protein discovery with energy rank alignment.
Sebastian Ibarraran, Shriram Chennakesavalu, Frank Hu, and Grant M. Rotskoff
ERA aligns learned sequence energies with experimental fitness rankings, enabling fast, data-efficient design loops. Use it to train geometric transformers, sample sequences, and map full fitness landscapes from limited data.
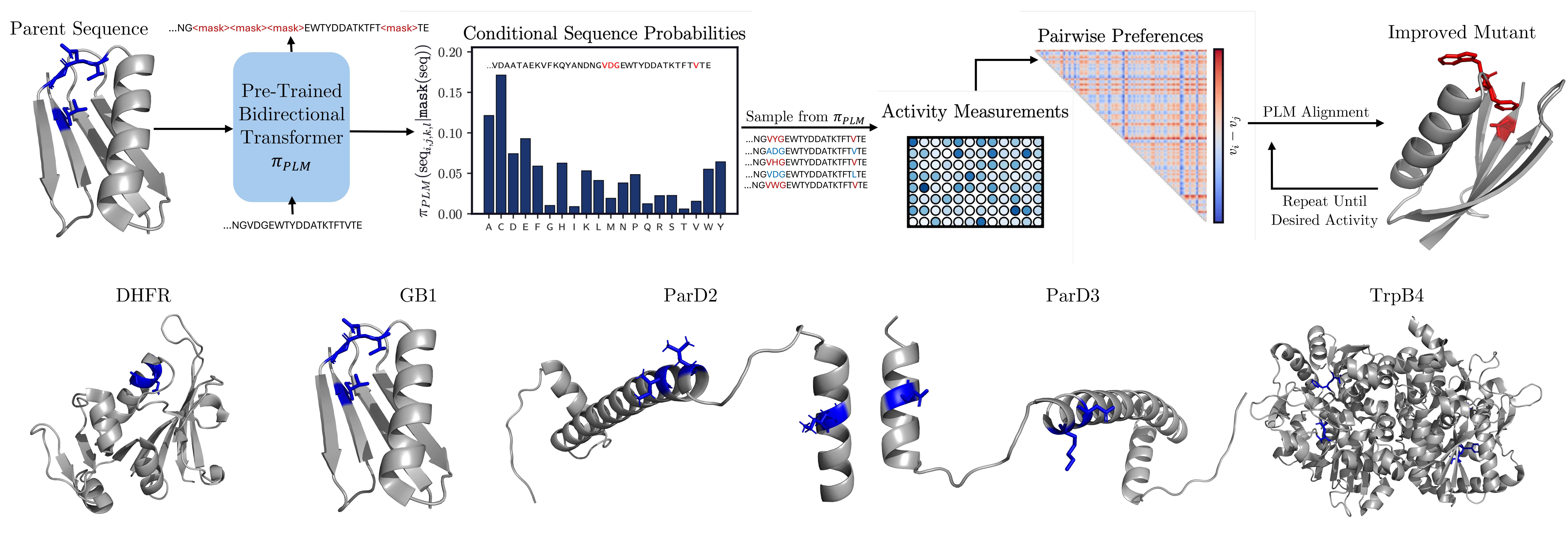
Preference Optimization for Few-Shot Experimental Design
| Method | DHFR | GB1 | ParD2 | ParD3 | TrpB4 |
|---|---|---|---|---|---|
| MLDE | 0.96 | 0.67 | 0.99 | 0.90 | 0.78 |
| ALDE | 0.96 | 0.80 | 0.99 | 0.99 | 0.89 |
| EVOLVEpro (ESM2) | 0.95 | 0.81 | 1.0 | 0.99 | 0.88 |
| EVOLVEpro (ESM3) | 1.0 | 0.77 | 1.0 | 0.99 | 0.79 |
| DPO | 1.0 | 0.78 | 1.0 | 0.99 | 0.81 |
| ERA | 1.0 ± 0.00 | 0.83 ± 0.04 | 1.0 ± 0.00 | 1.0 ± 0.00 | 0.91 ± 0.02 |
Energy rank alignment reframes few-shot protein engineering as preference optimization. Let $f(x)$ be a learned energy for sequence $x$. For any observed pair $(x_i, x_j)$ where $x_i$ is experimentally preferred to $x_j$, we enforce $f(x_i) < f(x_j)$ using a smooth pairwise loss. We optimize $$ \mathcal{L}_{\mathrm{ERA}}(\pi_\theta) = \mathbb{E}_{x \sim \mathcal{D},\, y,y' \sim \pi_{\mathrm{ref}}(\cdot \mid x)} \!\left[ D_{\mathrm{KL}}\!\left( p_\gamma(y \succ y') \,\|\, p_\theta \right) \right]. $$ which penalizes inversions while remaining stable under limited data. Optimizing this ranking loss aligns energies with relative fitness ordering instead of absolute calibration, improving generalization when only a handful of measurements are available.
Train using your data
Environment setup
uv sync
uv pip install -e .Train a model
pera_train \
train.data_root_path=./data \
train.data=GB1 \
train.trainer_args.max_epochs=100 \
train.trainer_args.devices=1Sample sequences
pera_sample \
infer.target=GB1 \
infer.num_samples=512 \
infer.network_filename=/path/to/checkpoint.ptCompute landscape
pera_compute_landscape \
compute_landscape.data=GB1 \
compute_landscape.data_root_path=./data